Investigating Harmful Algal Blooms in Ontario
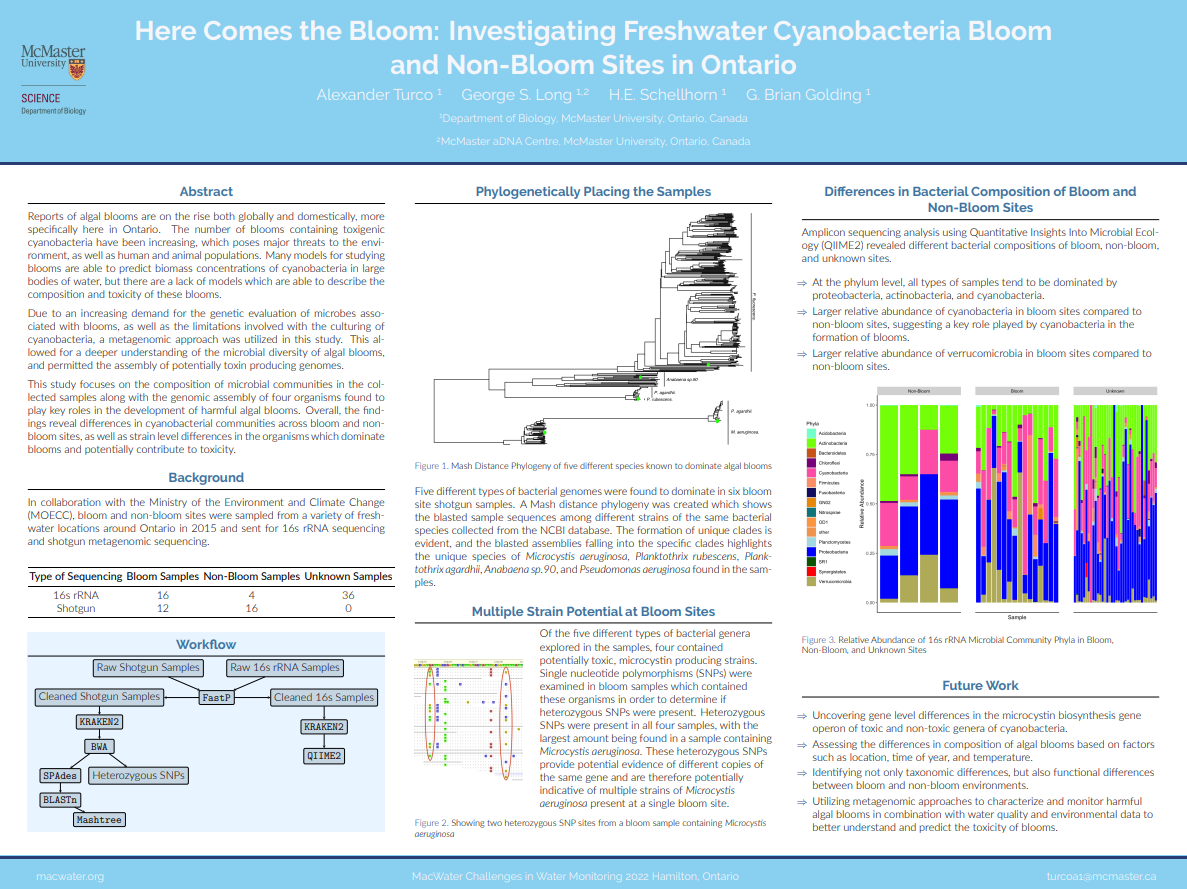
As a research assistant at McMaster University, I spent the summer exploring harmful algal bloom sites across Ontario. Under the supervision of Dr. Brian Golding and Dr. Herb Schellhorn, I conducted a metagenomic analysis of bloom and non-bloom sites using samples provided by the Ministry of Environment and Climate change. I examined the bacterial composition of samples, trimmed, merged, and assembled genomes of organisms known to contribute to the toxicity of blooms, and identified the potential for multiple strains of the same species to be present at a single bloom site. I created a poster to summarize some of the findings from this research. This poster was displayed at the MacWater (McMaster water group) challenges in water monitoring conference held on October 14 in Hamilton. Professors, graduate students, and those who work in industry could view and inquire about the poster and the work being done.
Open project